I will perform microbial rna seq dge analysis using deseq2 and snakemake
Biotechnology Researcher: Microbial Genomics and Bioinformatics
About this Gig
Welcome! As a PhD Researcher in Biotechnology & Microbiology, I provide expert-level analysis for Microbial RNA-Seq data.
I don't just run code; I am a published researcher who understands the science behind your results. My specialty is Differential Gene Expression (DGE) and identifying the key up/down-regulated genes for your study.
### What This Gig Offers (RNA-Seq DGE)
I offer three levels of analysis based on my proven pipeline (FastQC, HISAT2, featureCounts, DESeq2). (Please see my FAQ below regarding sample count!).
* Basic (The "Data"):
I will run the full pipeline up to Read Counts (Step 04). This package is perfect for researchers who use R/Python and just need the final Counts Matrix.
* Standard (The "Analysis"):
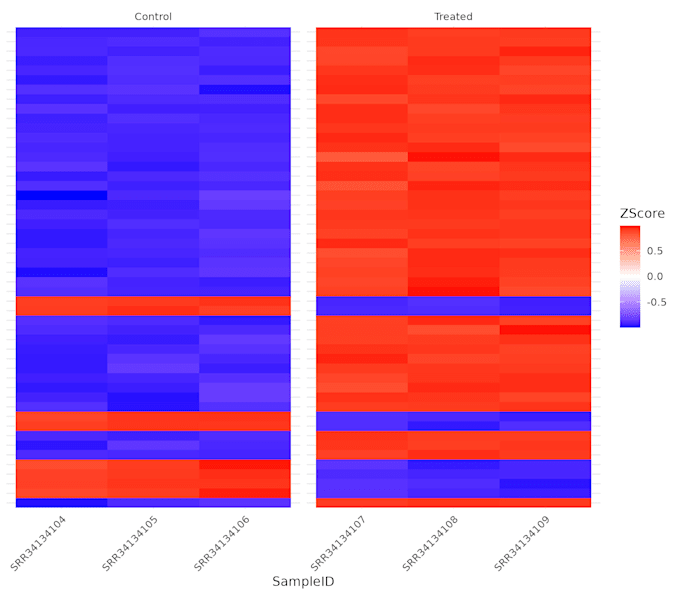
This is the full DGE Analysis (Step 05). You get everything in Basic, PLUS all the key visualizations: Volcano Plot, PCA Plot, & Heatmap.
* Premium (The "PhD Insight"):
The full publication-ready package. You get everything in Standard, PLUS: a full PDF report with my expert **Biological Interpretation** (the "why") AND the reproducible Jupyter Notebook used for your analysis.
**Please contact me before ordering to discuss your project.
Programming language:
Python
•
R
•
SPSS
Technology:
Jupyter Notebook
Expertise:
Cluster analysis
Tools:
Pandas
Other Data Analytics Services I Offer
FAQ
Your package prices don't mention the number of samples. Why?
The package prices ($30, $50, $80) are a baseline for small pilot projects (e.g., 1-3 samples). For larger cohorts (4, 6, or 20+ samples), please message me. We will discuss your goals, and I will send you a precise Custom Offer.
Why should I choose the Premium package? What's the value?
Premium is my PhD-level "Insight" service. I don't just provide a list of genes; I offer a comprehensive PDF report with **Biological Interpretation** (the "why"). You also get the **reproducible Jupyter script** for your publication.
What do you need from me to start?
I need 3 things: 1. Your raw FASTQ files (R1 & R2). 2. The reference genome (FASTA) and annotation (GFF/GTF) files. 3. Your metadata file (e.g., which samples are 'Control' vs 'Treated').



_hmj74d.jpg)